PLANT UPTAKE OF AMINO ACIDS AND PEPTIDES
By: Joshua R. Widhalm, Department of Horticulture and Landscape Architecture, Purdue University, United States
email: jwidhalm@purdue.edu

Amino acids are vital building blocks for synthesizing proteins in all organisms. In plants, free “proteinogenic” amino acids serve additional roles in the assimilation and transport of nitrogen, as signaling compounds, as osmolytes, and as precursors for making various hormones, cofactors, and other major compounds like chlorophyll. Plants also use amino acids to collectively produce thousands of specialized compounds that facilitate ecological interactions and provide adaptive responses to environmental stresses.
Amino acids in plants can also be bound in polymeric forms as small peptides, which are typically comprised of 5 to 60-amino acids. Certain small peptides serve hormone-like roles in plant growth and development (Roy et al., 2018). Others function in response to wounding, pathogen infection, nutrient imbalance, drought, or high salinity (Chen et al., 2020). While many endogenously produced small peptides are gene-encoded, some are produced through cleavage of preproteins and posttranslational modifications (Matsubayashi, 2014). Externally applied peptides as components of protein hydrolysates or those produced synthetically with random sequences have also been found to elicit biological activities (Roy et al., 2018). However, understanding of the molecular mechanisms underlying how free amino acids and their polymeric forms—as peptides and proteins—are taken up from the environment has only recently started to emerge.
Investigations into the biochemical processes mediating uptake and transport of amino acids and peptides started in the 1980s and 1990s with uptake assays using membrane vesicles isolated from various plant tissues. Using radioactively labeled substrates and fluorescent labeling methods, the occurrence of multiple amino acid and peptide transport systems was revealed (Tegeder and Rentsch, 2010). Since then, members from at least a dozen transporter families have been identified in the nonmycorrhizal model species Arabidopsis thaliana. Functional studies—using a combination of heterologous expression systems, gene knockout and overexpression lines, and expression analyses to determine organ/tissue localization—have revealed roles for transporters in soil amino acid uptake, cellular import and export, and/or long-distance transport throughout the plant vascular system (Tegeder, 2014).
Plasma membrane-localized transporters involved in root amino acid uptake belong to three families within the Amino acid-Polyamine-Choline (APC) transporter superfamily: (i) the Amino Acid Permeases (AAPs), which function as broad substrate transporters with moderate affinities; (ii) the Lysine/Histidine-like Transporters (LHTs), which import neutral and acidic amino acids; and (iii) the Proline and Glycine Betaine Transporters (ProTs), which transport the osmolytes proline and glycine betaine (Tegeder and Masclaux-Daubresse, 2018).

AtPTR1, a member of the peptide transporter/nitrate transporter 1 (PTR) family is involved in uptake of dipeptides into roots (Komarova et al., 2008) (Figure 1). Beyond that, little else is known root peptide uptake, although other PTRs and members of the ATP Binding Cassette (ABC) transporter and Oligopeptide Transporter (OPT) families are likely involved (Tegeder and Rentsch, 2010; Roy et al., 2018).
In mycorrhizal plants, fungal symbionts contribute to the uptake of amino acids and peptides. Mycorrhizas also exhibit proteolytic activities that, together with those of free-living microbes in the soil, make amino acids bound in proteins and peptides available to plants (Näsholm et al., 2009) (Figure 1).
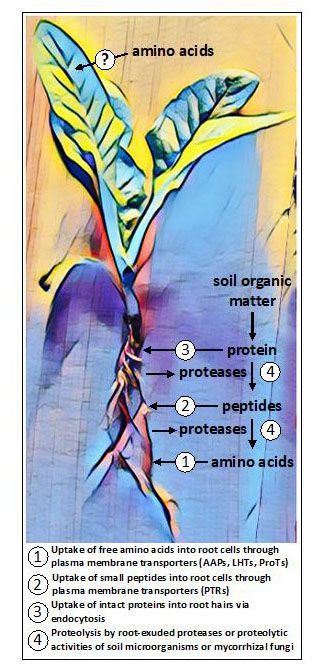
Figure 1. Potential routes of amino acid uptake collectively found in plants. In roots, amino acids can be taken up in their free (monomeric) forms or in polymeric forms as small peptides or proteins. Some plants release proteases to convert proteins and peptides in the soil into free amino acids, while other plants rely on the activities of fungal symbionts for proteolysis and uptake. Emerging evidence is indicating that certain plants may be able to acquire amino acids exogenously applied to aerial tissues, although the presumptive routes of uptake are unknown.
It was only more recently discovered that roots of several distinct nonmycorrhizal plants exude proteolytic enzymes belonging to the cysteine protease family (Adamczyk et al., 2010) that can break down proteins at the root surface and in the apoplast of the root cortex (Paungfoo-Lonhienne et al., 2008). The released small peptides and free amino acids are then available for uptake by plasma membrane-localized transporters. Plant cell membranes also contain leucine-rich repeat receptor kinases (LRR-RKs) that sense peptide and protein ligands (Hohmann et al., 2017). These receptors form the first layer of the plant immune system and members expressed in roots help to control formation of diffusion barriers in the Casparian strip (Okuda et al., 2020). Intact proteins are also reportedly able to enter root hairs by endocytosis and are then digested inside plant cells (Paungfoo-Lonhienne et al., 2008).
In the soil, plants are easily outcompeted by microorganisms for organic nitrogen. Microbes have faster turnover rates and larger surface area to volume ratios, giving them increased access to substrates in the soil. This then raises questions about the degree to which amino acids, peptides, and proteins serve as significant sources of nitrogen for plants (Näsholm et al., 2009). As exogenously applied amino acids, peptides, and proteins have beneficial effects (Colla et al., 2017b) beyond nitrogen metabolism, this may point to other roles for amino acids and peptides taken up from soil.
While much research has focused on the acquisition of amino acids from soil, some plants are capable of taking them up via aerial tissues. In the pitfall traps of carnivorous plants like Nepenthes species, for example, nutrients released from digested prey, including amino acids, peptides, and proteins, are recovered by the plant. Following digestion, the molecules penetrate the cuticle via pores and enter the apoplast where they are endocytosed into cells, and then translocated toward the mesophyll and further incorporated into metabolism (Adlassnig et al., 2012).
Epiphytic plants (also known as “air plants”) primarily obtain moisture and nutrients from the air and other non-soil debris around them. These non-parasitic plants grow on top of other plants, and include many types of mosses, ferns, orchids, and bromeliads. Epiphytic bryophytes, which do not have cuticle barriers and lack developed root and vascular systems, were shown by 15N-labeling studies to take up glycine through foliar application (Song et al., 2016). Epiphytic bromeliads, like “Spanish moss” (Tillandsia usneoides) and the “potbelly airplant” (T. paucifolia), are non-parasitic monocots with reduced vascular systems that use their roots to attach to nearby plants and then gather nutrients, including amino acids, via specialized leaf trichomes (Nyman et al., 1987).
What about other types of plants? Tracer studies with 15N-labeled amino acids showed that creeping bentgrass (Stiegler et al., 2013) and peach tree (Furuya and Umemiya, 2002) leaves are capable of absorbing amino acids through their foliage. A recent study by McCoy et al. (McCoy et al., 2020) further demonstrated that 15N,13C double-labeled glutamate exogenously applied to creeping bentgrass foliage is incorporated into proline and γ-aminobutyric acid (GABA). This work is significant because it suggests that mineralization on the leaf surface is a minor fate for applied amino acids and demonstrates that exogenously supplied amino acids can be incorporated into plant metabolism.

Foliar application of protein hydrolysates has been observed to increase productivity, environmental stress tolerance, and other physiological characteristics in a number of agriculturally-relevant species, including tomato (Colla et al., 2017a), fruit trees (Tanou et al., 2017), and soybean (Teixeira et al., 2018). While applications often contain surfactants to increase permeance of hydrophilic amino acids and small peptides across the hydrophobic cuticle, other factors, such as climate (Pecha et al., 2012) and the presence of aqueous pores (Schönherr, 2006) also affect uptake.
Once amino acids and peptides cross the cuticle barrier, they enter the cell wall space (apoplast) where they can be taken up by one or more plasma membrane-localized transporters. From here, they can also—a priori—bind LRR-RKs or be transported to other parts of the plant. The most likely route of long-distance transport is via the apoplastic phloem pathway, which imports amino acids from the apoplast into the phloem (Tegeder and Masclaux-Daubresse, 2018). The transporters involved include members from the same AAP class contributing to root uptake of amino acids from soil (Santiago and Tegeder, 2016). Interesting to note, operation of the apoplastic phloem pathway in plants typically relies on the export of amino acids from leaf source cells into the apoplast. Thus, it can be envisioned that exogenously supplying amino acids through the foliage in effect bypasses this step, thereby influencing aspects of fruit quality and root architecture seen when applying protein hydrolysate-based biostimulants (Colla et al., 2015).



